Computational Medicine
UCLA Health technical project manager Robert Smith joins for a discussion on the state of personalized medicine and computational medicine.
The goal of personal genomics is to move toward being able to interpret each person’s individual genetic sequence, and predict it’s effect on their bodies and health.
Preoperative predictions of in-hospital mortality using electronic medical record data | bioRxiv
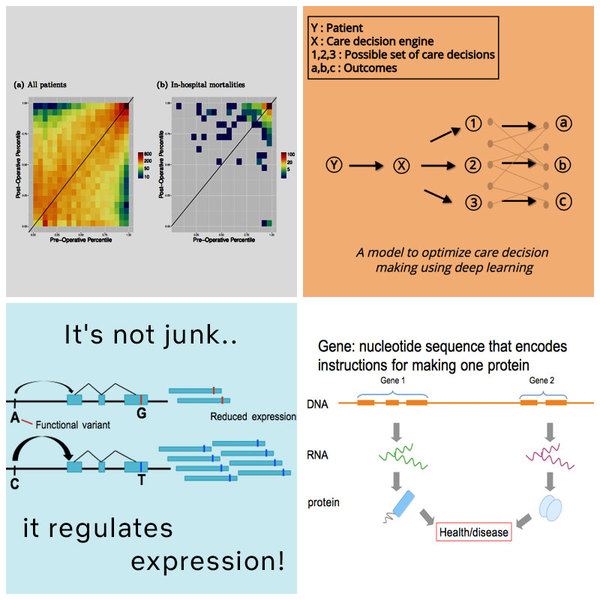
Predicting preoperative in-hospital mortality using readily-available electronic medical record (EMR) data can aid clinicians in accurately and rapidly determining surgical risk. While previous work has shown that the American Society of Anesthesiologists (ASA) Physical Status Classification is a useful, though subjective, feature for predicting surgical outcomes, obtaining this classification requires a clinician to review the patient’s medical records. Our goal here is to create an improved risk score using electronic medical records and demonstrate its utility in predicting in-hospital mortality without requiring clinician-derived ASA scores.
Large-scale transcriptome-wide association study identifies new prostate cancer risk regions | Nature Communications
Although genome-wide association studies (GWAS) for prostate cancer (PrCa) have identified more than 100 risk regions, most of the risk genes at these regions remain largely unknown. Here we integrate the largest PrCa GWAS (N = 142,392) with gene expression measured in 45 tissues (N = 4458), including normal and tumor prostate, to perform a multi-tissue transcriptome-wide association study (TWAS) for PrCa. We identify 217 genes at 84 independent 1 Mb regions associated with PrCa risk, 9 of which are regions with no genome-wide significant SNP within 2 Mb. 23 genes are significant in TWAS only for alternative splicing models in prostate tumor thus supporting the hypothesis of splicing driving risk for continued oncogenesis. Finally, we use a Bayesian probabilistic approach to estimate credible sets of genes containing the causal gene at a pre-defined level; this reduced the list of 217 associations to 109 genes in the 90% credible set. Overall, our findings highlight the power of integrating expression with PrCa GWAS to identify novel risk loci and prioritize putative causal genes at known risk loci.

